I’ve been working on a project intermittently for some time, and I recently packaged it up and published it on GitHub, hoping it could be useful to


I’ve been working on a project intermittently for some time, and I recently packaged it up and published it on GitHub, hoping it could be useful to
In a previous blogpost: (Generating Unusual Molecules with Genetic Algorithms), I showcased the propensity of a genetic algorithm to generate unusual molecules. This illustrates that this generative

We’ve just updated Scikit-Mol[1] to version 0.3.0. Scikit-Mol was covered in a previous blogpost. The big news in this update is the support for pandas output (and
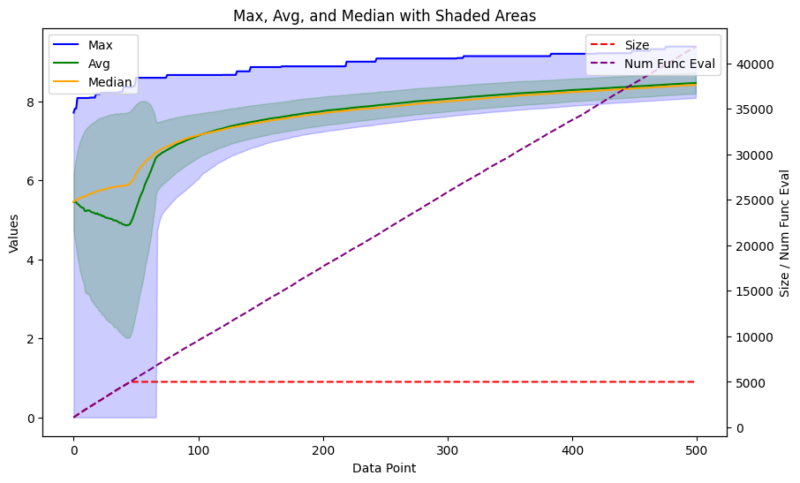
I’ve long been working with generative models, mostly centered around SMILES-based deep learning models. However, I’ve been wanting to try out genetic algorithms for some time. Using

I’d like to share a post about a project I’ve been involved in developing—Scikit-Mol. I believe it’s a noteworthy project deserving attention. It has already been featured
We’ve known since 2016 that LSTM networks can be used to generate novel and valid SMILES strings of novel molecules after being trained on a dataset of
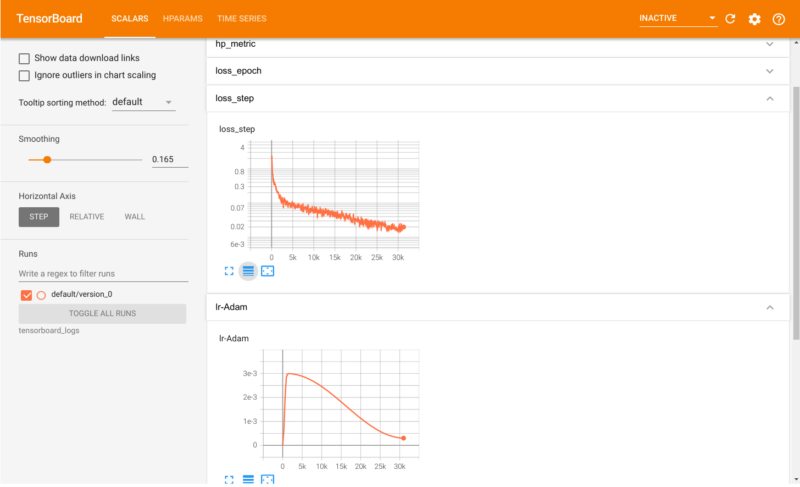
In the last blogpost I covered how LSTM-to-LSTM networks could be used to “translate” reactants into products of chemical reactions. Performance was however not very good of
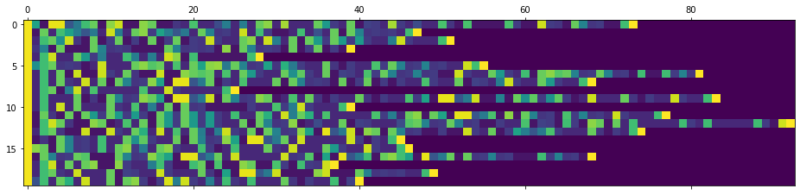
In this blogpost I’ll show how to predict chemical reactions with a sequence to sequence network based on LSTM cells. It’s the same principle as IBM’s RXN

I have been writing a lot about how to use SMILES together with deep learning architectures such as RNNs and LSTM networks to perform various cheminformatic and
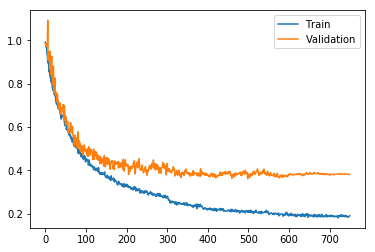
Last blog-post I showed how to use PyTorch to build a feed forward neural network model for molecular property prediction (QSAR: Quantitative structure-activity relationship). RDKit was used